Illumina sequencing submission
Illumina Short Read Sequencing Technology
Short-read sequencing, primarily powered by Illumina technology, is one of the most widely used methods for generating high-throughput genomic data. Illumina sequencing platforms utilize sequencing by synthesis (SBS), a method that involves the stepwise incorporation of fluorescently labeled nucleotides during DNA synthesis. This process enables massively parallel sequencing, producing millions to billions of short reads (typically 50–300 base pairs) per run.
The workflow typically involves library preparation, where DNA or RNA is fragmented and adapters are ligated to form a final library molecule (structure illustrated below).
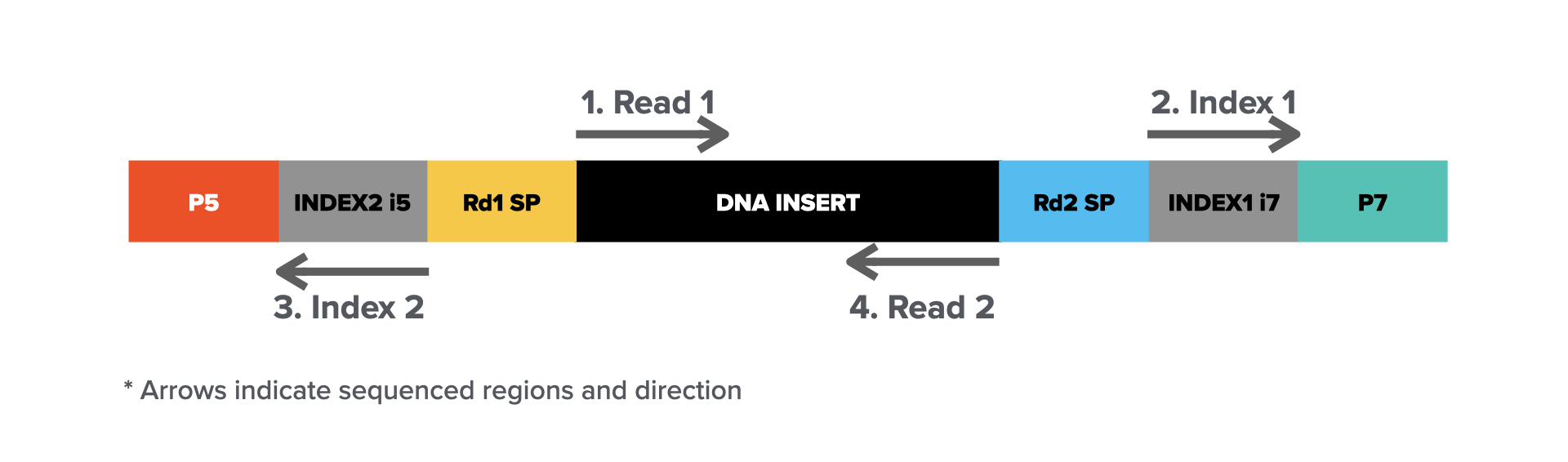
Sequencing starts by cluster amplification on a flow cell, cyclic sequencing-by-synthesis (steps illustrated above), and base calling via imaging. The high accuracy, scalability, and cost-effectiveness of Illumina sequencing make it a standard for applications such as whole-genome sequencing (WGS), RNA sequencing (RNA-seq), exome sequencing, and epigenomic studies. However, the short read length presents challenges in resolving complex genomic regions, structural variations, and long-range connectivity, which are areas where long-read sequencing technologies complement short-read approaches.
Multiplexing for High-throughput Sequencing
Multiplexing in Illumina sequencing allows multiple samples to be pooled and sequenced in a single run by incorporating unique index sequences (barcodes) into each library. These indexes are added during library preparation and later used to demultiplex reads after sequencing. Multiplexing improves efficiency, reduces costs, and enables comparative studies by maximizing data output per run. Dual-indexing helps prevent errors like index hopping, ensuring accurate sample identification. Proper library balancing and careful index selection are crucial for high-quality results in applications like RNA-seq, metagenomics, and whole-genome sequencing. If you have any questions, please contact seq-team@ny.czbiohub.org.
Resources
- Illumina Website
- Next-Generation Sequencing Basics
- Illumina Adapter Sequences
- Indexed Adapter Pooling Guide
- Sequencing Coverage Calculator
- Adapter Dimer Problem
- Removing Adapter Dimers with SPRI Beads
Sequencing submission overview
This sequencing submission system is for Biohub groups and onboarded Biohub Investigators only. If you are a new investigator, please reach out to Pablo Garcia-Nieto with a brief description of your interests and proposed scope of work for sequencing-based experiments. We currently only accept prepared and pooled libraries. If you have questions about what library protocol to use based on your application of interest please refer to Illumina's library preparation kits or reach out to our team at seq-team@ny.czbiohub.org.
Submission instructions
Please read the current submission instructions carefully before proceeding with a submission.
Use the submission form to create an initial submission for Illumina Sequencing. Please note that in this system you will need to download, complete and upload a samplesheet as a CSV file during the initial submission process, prior to dropping off your library pool. Please refer to the instructions for sample sheet requirements.
Library QC
Once we receive your library we will perform a quality check before loading the library on the sequencer. We will reach out to the submitter via email if we need further information.
Sequencing turnaround time
The turnaround time for sequencing is dependent on the number of requests in our queue. Expect a 2-3 week turnaround time from full submission (form completion and library drop-off) to data delivery, expect longer turnaround for partial flowcell requests and wrong sample sheets provided.
Data delivery
The demultiplexed data will be delivered to you via email, which contains instructions to access your data. For external collaborators, you will receive an AWS token for data download from the cloud. Please make sure that you have AWS CLI installed on your computer. The token is valid for 36 hours, so please download your data as soon as you receive an email from the Genomics Platform team containing the token.
Sample submission form
Once you have read the instructions above, please click the button below to proceed to the sample submission form.
Proceed to the submission form